 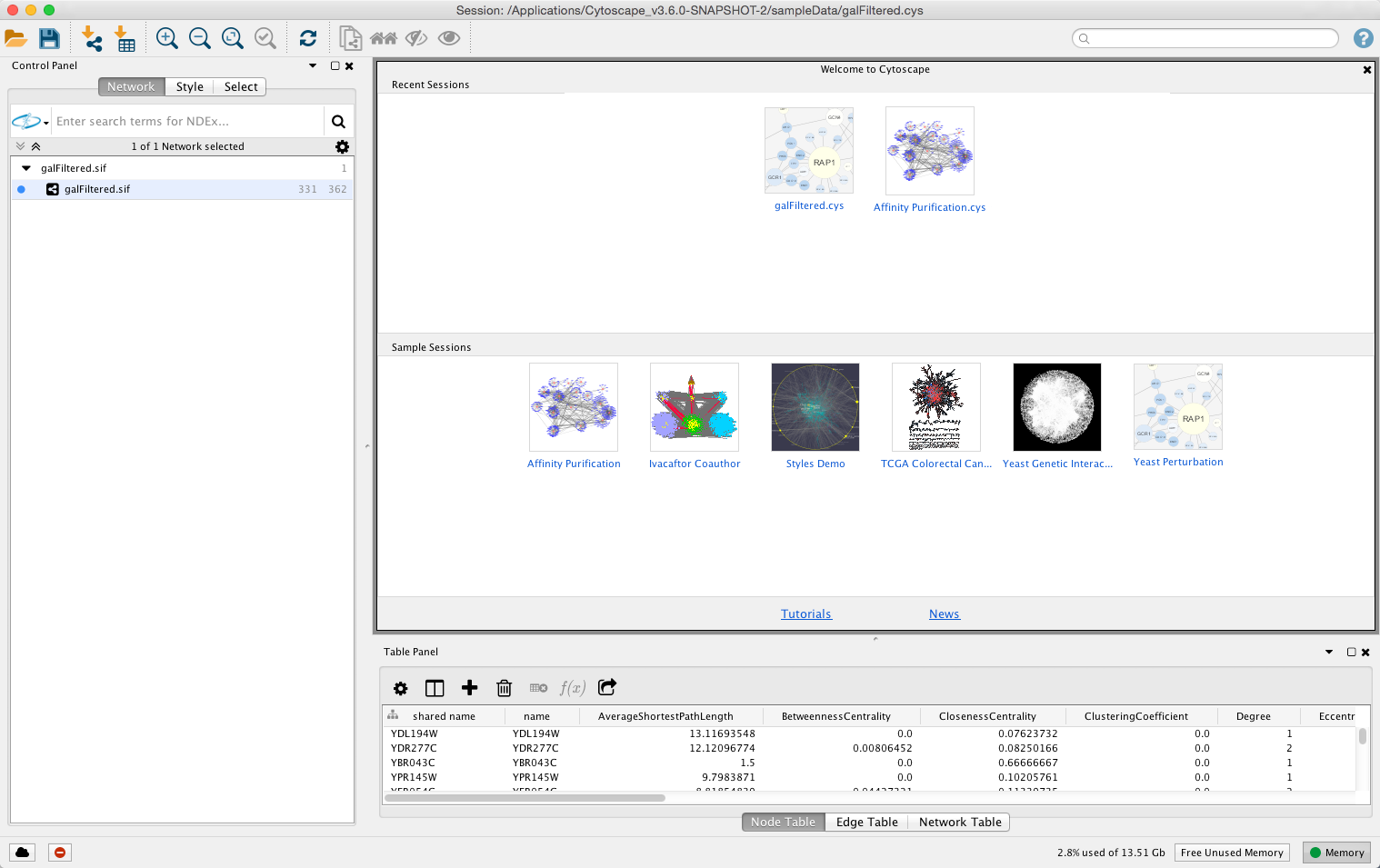
N.B.: this is not a problem on Linux! (If you have a 64-bit version of Windows and are running a 64-bit version of Java, you can also try using this Cytoscape.vmoptions file. into Cytoscape using semi-automated methods, including Linux-based scripts. : A non-minified UMD build with all dependencies included in the bundle. You typically need to use a programmers editor and make sure to set the line-end configuration to line-feeds only! Another option on Windows is to create the Cytoscape.vmoptions file on Mac OS or Linux and then to copy it into the Windows installation directory. Cytoscape is a bioinformatic data analysis and visualization platform that. Allow selection of multiple networks in network list panel ( CytoPanel 1). Also, some F keys interfere with normal Mac operation. on the Mac, the command key instead of the ctrl key should be used. This is difficult to accomplish on Windows on which the default is line termination with carriage-return/line-feed pairs. Currently Cytoscape shortcuts are more suited for Windows and Linux and don't work very well on the Mac. a "newline" character) and must not contain any carriage return characters. The problem with Windows is that the lines need to be terminated with a single line feed character (a.k.a. Excel spreadsheet download - Cytoscape for Linux 3.9. Just like with Option A, you can use the Cytoscape.vmoptions file. Option B: Using the Cytoscape icon (Windows and/or Linux systems) This file allocates a maximum of 5GiB of heap memory and 10MiB of stack space per thread. Cytoscape is an open source software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data. The exercises will teach you how to : retrieve networks for proteins or small-molecule compounds of interest retrieve networks for a disease or an arbitrary topics in PubMed layout and visually style the resulting networks. A typical Cytoscape.vmoptions file would look like this: -Xmx5G In these exercises, we will use the stringApp for Cytoscape to retrieve molecular networks from the STRING and STITCH databases. Cytoscape is a Java application verified to run on Linux, Windows. Other options you may use may depend on the particular JVM that you are running. The central organizing metaphor of Cytoscape is a network (graph), with genes. 
You can add an "M" for megabytes or a "G" for gigabytes. The most popular and important option is "-Xmx" followed immediately (no spaces!) by the amount of heap memory. Option B: Using the Cytoscape icon (Windows and/or Linux systems). Modify section 6.4.2.1.1 Climatic Data Design Conditions to reference ASHRAE 169 and as a national standard reference for climatic data.Winter design temperatures are based upon the mean extreme annual temperature and summer conditions are the 1 percent annual cooling design conditions. Make sure to add exactly one JVM option per line and that the last line has a line break at the end. from Notes on memory consumption, Cytoscape User Manual. Option A: Command line startup (All operating systems/platforms)Ĭreate a Cytoscape.vmoptions file in the. All of them will change Cytoscape's default memory parameters except starting from the command line. Note: Windows builtin zip cannot uncompress this archive, please use a 3rd party utility like WinZip. There are a number of ways to change Cytoscape's memory allocation, depending on your preferred method of opening the application. Bioinformatics, 27(3), 431-432.Changing memory allocations on Windows, Mac, and Linux machines Cytoscape 2.8: new features for data integration and network visualization. Although Cytoscape was originally designed for biological research, now it is a general platform for complex network. Cytoscape: software for visualization and analysis of biological networks. Cytoscape is an open source software platform for visualizing molecular interaction networks and biological pathways and integrating these networks with annotations, gene expression profiles and other state data. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Shannon, P., Markiel, A., Ozier, O., Baliga, N.You can look for Cytoscape in the Ubuntu start menu or run it via terminal using the following command: $ sudo chmod u+x Cytoscape_3_7_2_unix.sh Running Cytoscape Provide access permission to the downloaded installer. $ sudo apt-get upgrade Downloading CytoscapeĬhange to the directory where you wish to download the software. Open the terminal (Ctrl+T) and type the following commands: Let’s update and upgrade the system first. In this article, we will install Cytoscape on Ubuntu. Cytoscape is a software for the easy visualization of complex networks. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
